D. Bhargava*, A. Chowdhury*, and D. H. Dube. "Chemical tools to study and modulate glycan-mediated host-bacteria interactions." Current Opinion in Chemical Biology, 2025, 87, https://doi.org/10.1016/j.cbpa.2025.102603
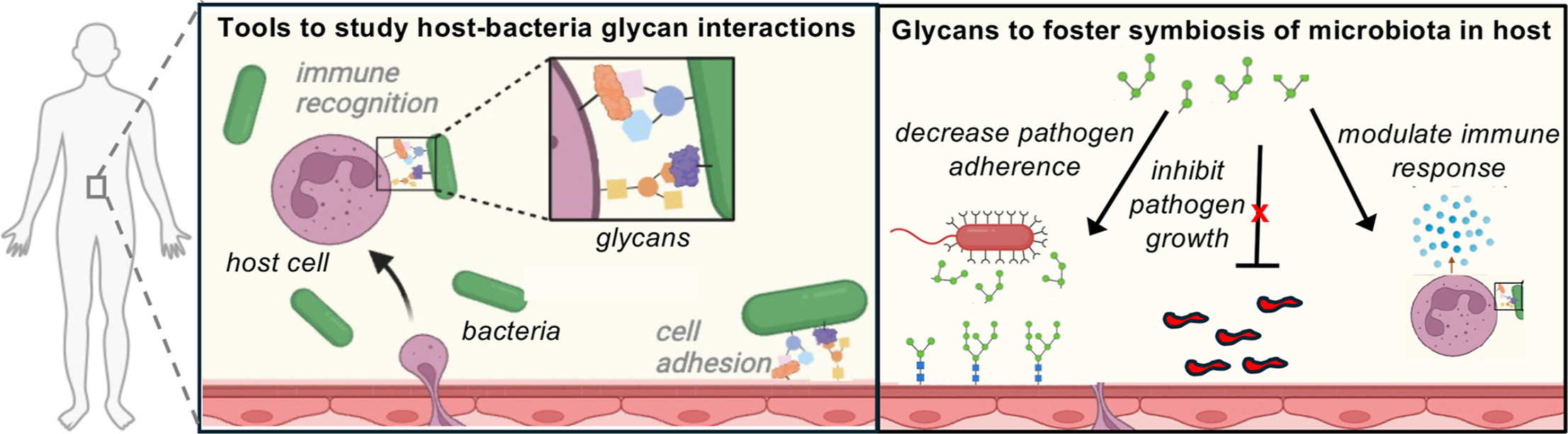
K. A. Barrett*, F. J. Kassama*, W. Surks*, A. Mulholland*, K. D. Moulton, and D. H. Dube. "Helicobacter pylori glycan biosynthesis modulates host recognition and response." Fronteirs in Cellular and Infection Microbiology, 2024, 14, https://doi.org/10.3389/fcimb.2024.1377077

D. Calles-Garcia and D. H. Dube. "Chemical biology tools to probe bacterial glycans." Current Opinion in Chemical Biology, 2024, 80, https://doi.org/10.1016/j.cbpa.2024.102453


I. Quintana,* A. Paul, A. Chowdhury,* K. D. Moulton, S. S. Kulkarni, and D. H. Dube. "Thioglycosides act as metabolic inhibitors of bacterial glycan biosynthesis." ACS Infectious Diseases., 2023, 9, 2025-2035. https://doi.org/10.1021/acsinfecdis.3c00324
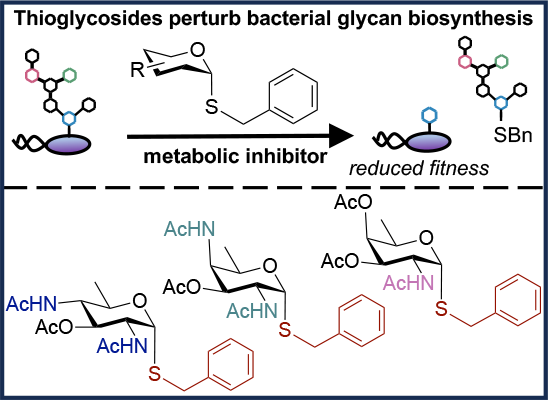

K. Barrett* and D. H. Dube. "Chemical tools to study bacterial glycans: a tale from discovery of glycooproteins to disruption of their function." Israel J. Chem., 2022, https://doi.org/10.1002/ijch.202200050
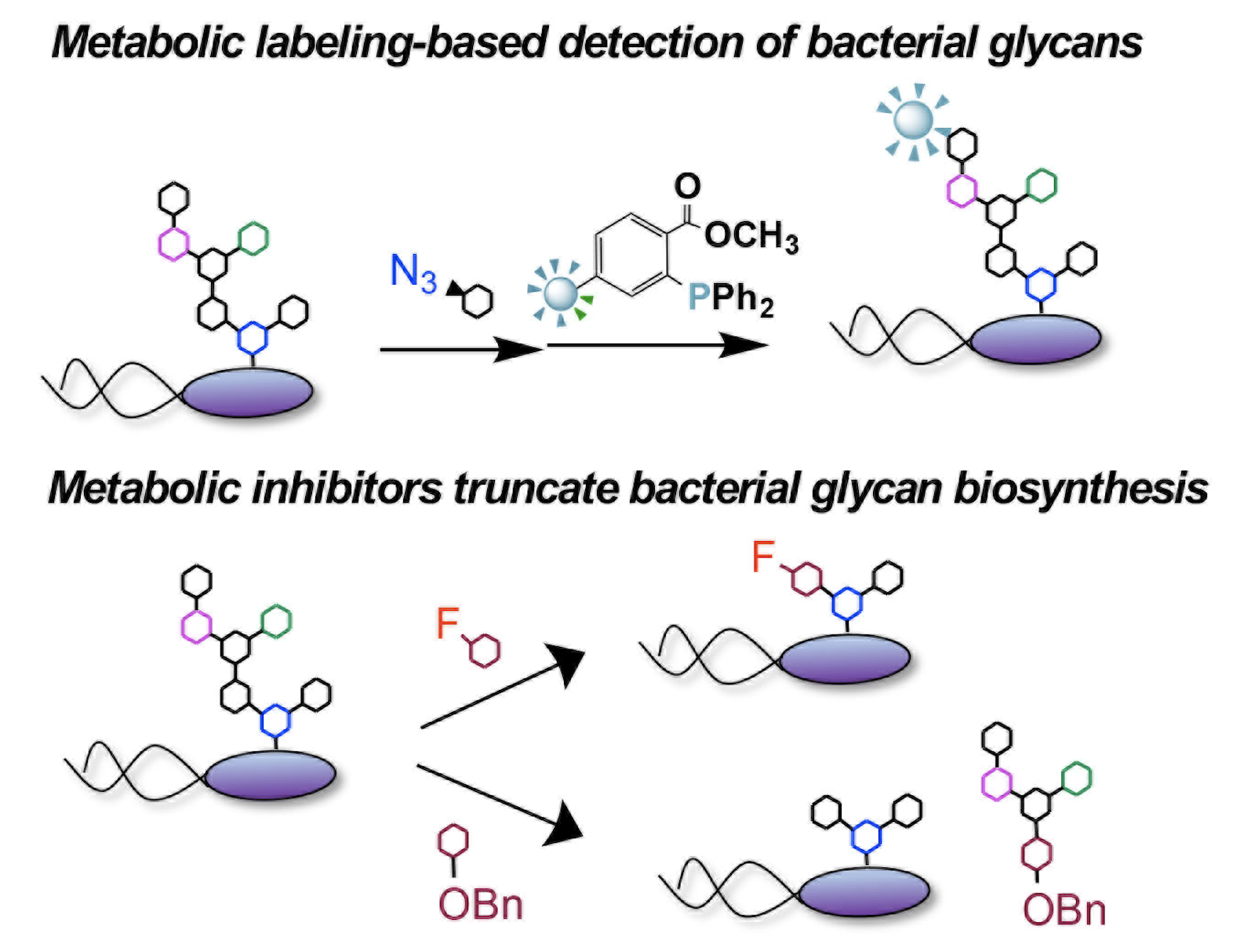
P. Luong,* A. Ghosh, K. D. Moulton, S. S. Kulkarni, and D. H. Dube. "Synthesis and application of rare deoxy amino L-sugar analogs to probe glycans in pathogenic bacteria." ACS Infectious Diseases., 2022, 8, 889-900.
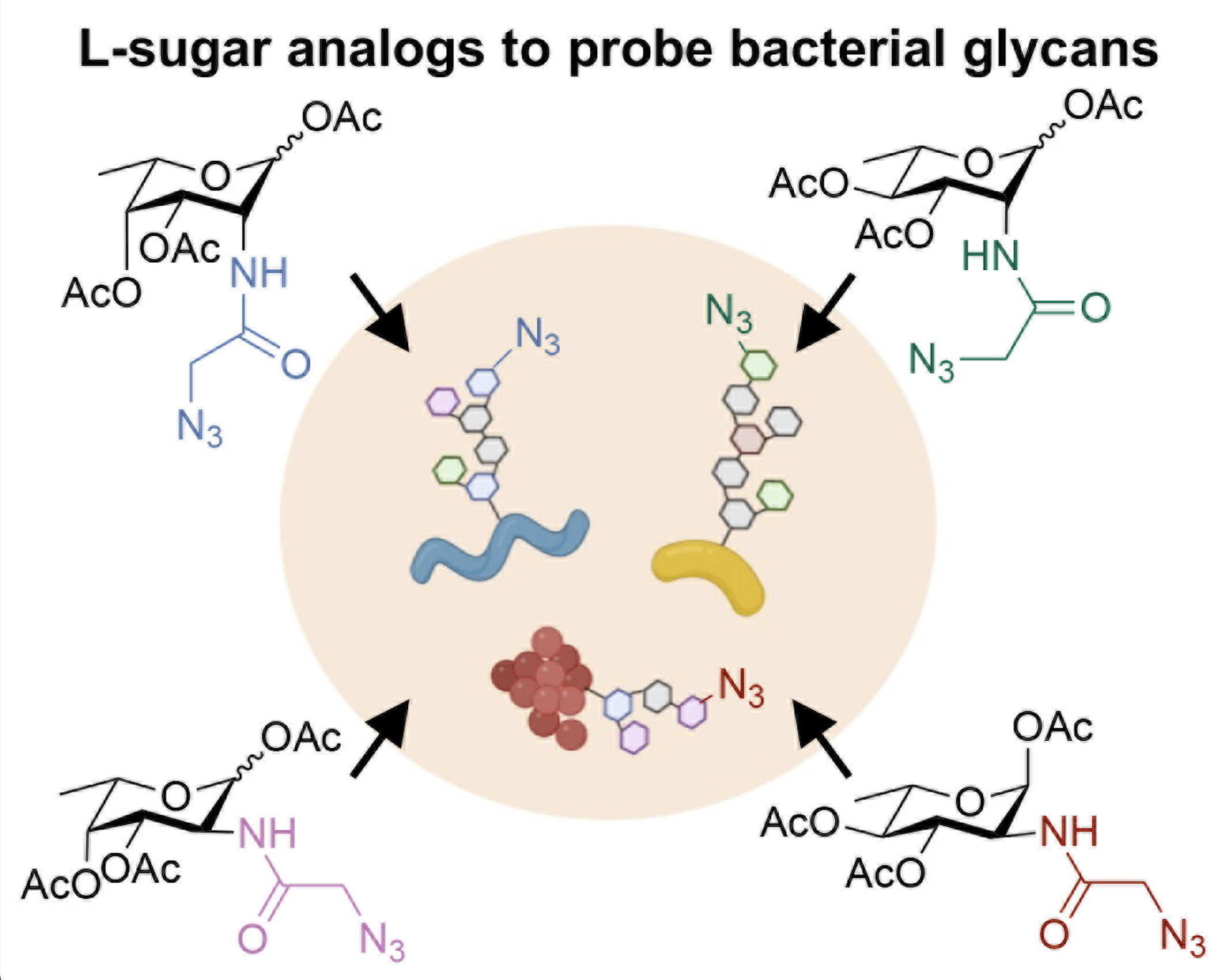
P. Luong* and D. H. Dube. "Dismantling the bacterial glycocalyx: chemical tools to probe, perturb, and image bacterial glycans." Bioorg. Med. Chem., 2021, 42, 116268.
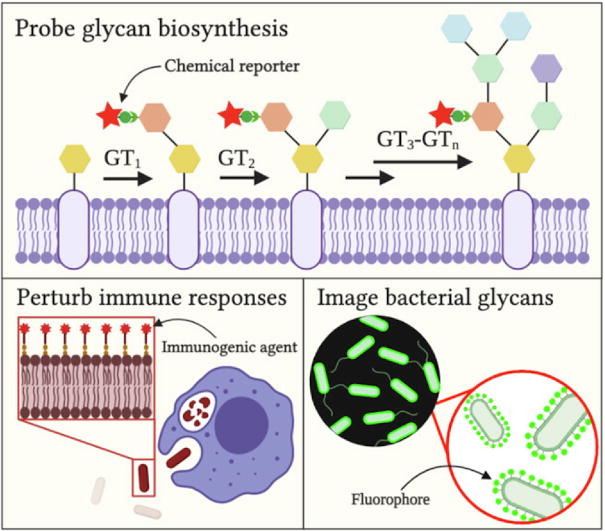
K. D. Moulton, A. P. Adewale*, H. A. Carol*, S. A. Mikami*, and D. H. Dube. "Metabolic glycan labeling-based screen to identify bacterial glycosylation genes." ACS Infectious Diseases, 2020, 6, 3247-3259.
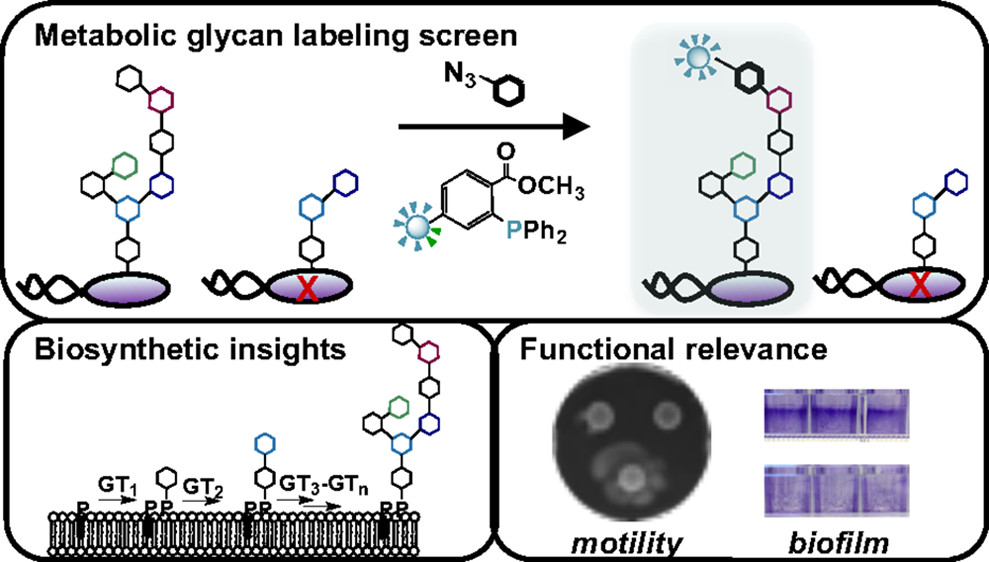
D. A. Williams*†, K. Pradhan†, A. Paul, I. R. Olin*, O. T. Tuck*, K. D. Moulton, S. S. Kulkarni, and D. H. Dube. "Metabolic inhibitors of bacterial glycan biosynthesis." Chemical Science, 2020, 11, 1761-1774.
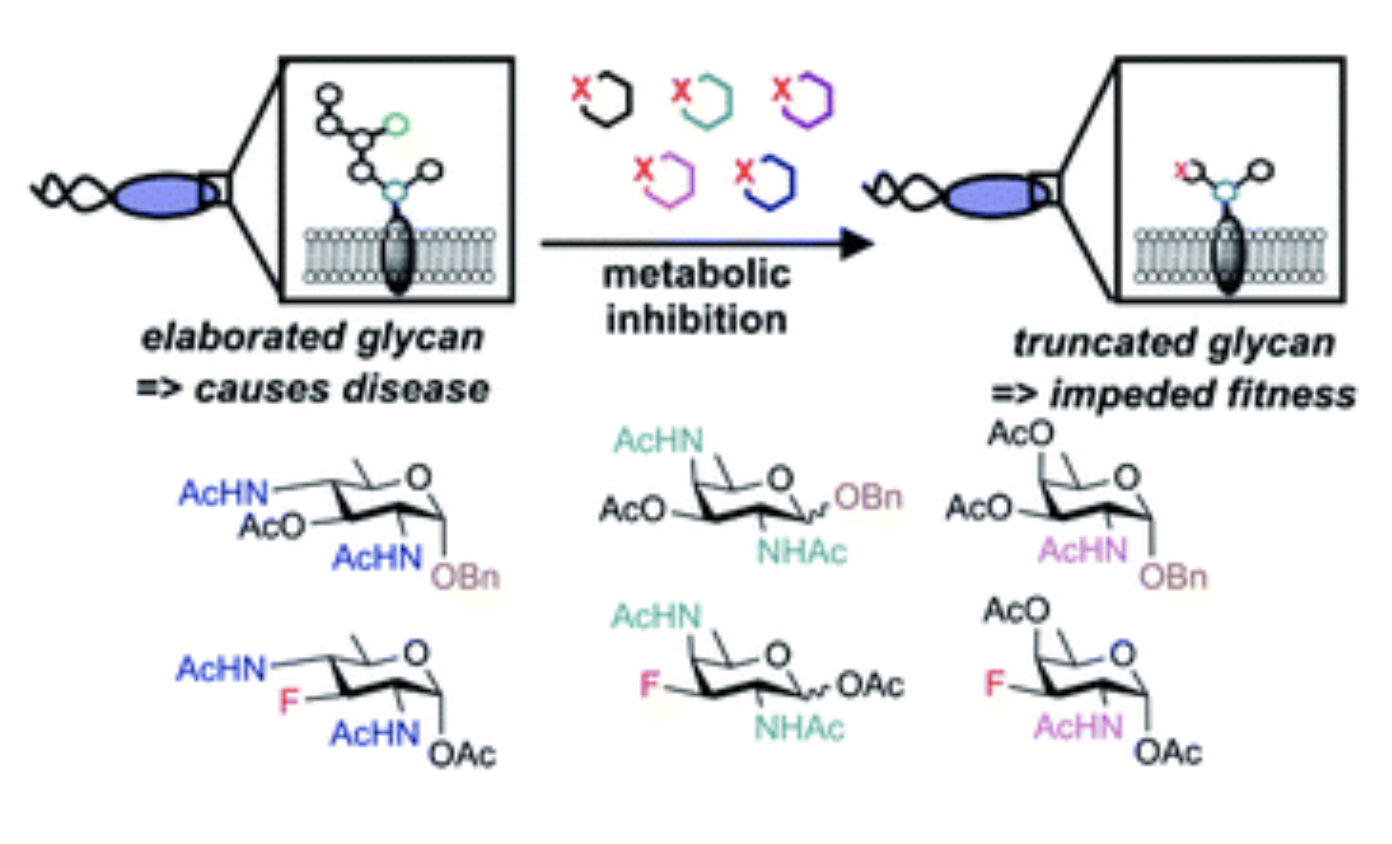
D. H. Dube, D. A. Williams*, "Metabolic glycan engineering in live animals: using bioorthogonal chemistry to alter cell surface glycans." In Handbook of in vivo chemistry in mice: from flask to animal, K. Tanaka, K. Vong eds., Wiley-VCH (Germany), 2020, https://doi.org/10.1002/9783527344406.ch8
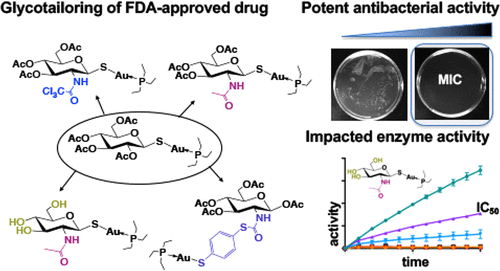
T. D. Epstein*†, B. Wu†, K. D. Moulton, M. Yan, D. H. Dube, "Sugar-modified auranofin analogs are potent inhibitors of the gastric pathogen Helicobacter pylori" ACS Infectious Diseases, 2019, 5, 1682-1687.
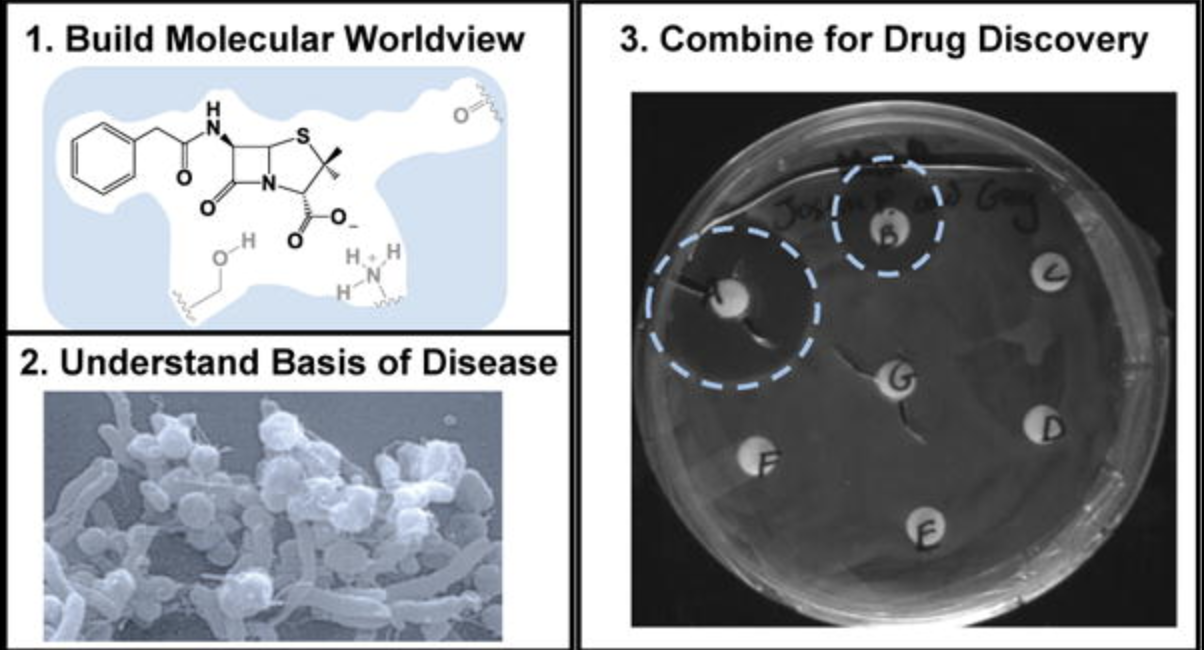
D. H. Dube, "Design of a drug discovery course for non-science majors" Biochem. Mol. Biol. Educ., 2018, 46, 327-335.
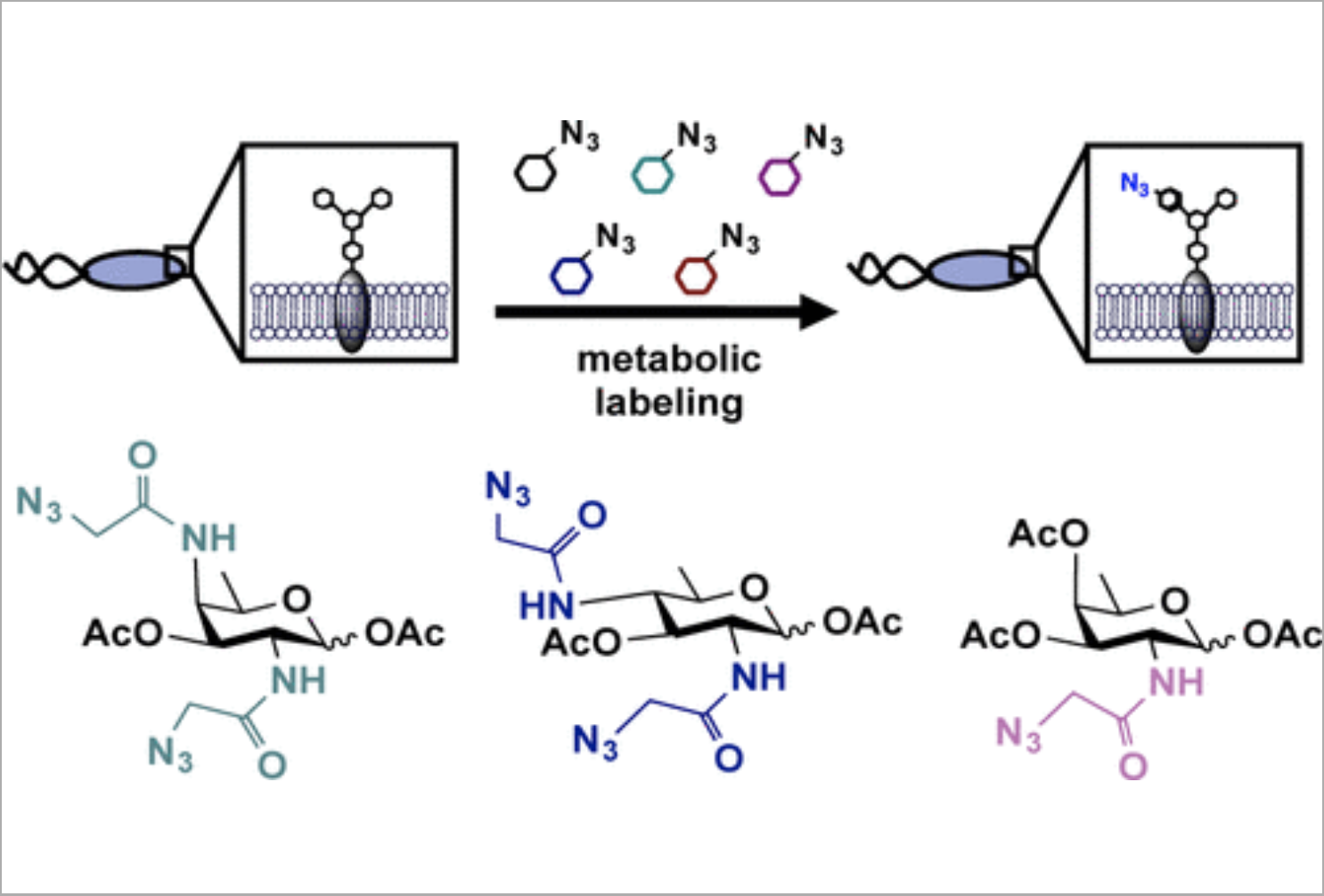
E. L. Clark*, M. Emmadi, K. L. Krupp*, A. R. Podilapu, J. D. Helble*, S. S. Kulkarni, and D. H. Dube, "Development of rare bacterial monosaccharide analogs for metabolic glycan labeling in pathogenic bacteria" ACS Chem. Biol., 2016, 11, 3365-3373.
K. R. Farnham and D. H. Dube, "A semester-long project-oriented biochemistry laboratory based on Helicobacter pylori urease" Biochem. Mol. Biol. Educ., 2015, 43, 333-340.
V. Tra* and D. H. Dube, "Chemical tools to detect and target Helicobacter pylori's glycoproteins" In Glycoscience: Biology and Medicine, T. Endo, P. Seeberger, G. Hart, C-H Wong, N. Taniguchi, ed., Springer (Japan), 2014.
V. Tra* and D. H. Dube, "Glycans in pathogenic bacteria -- potential for targeted covalent therapeutics and imaging agents," Chem. Commun., 2014, 50, 4659-4673.
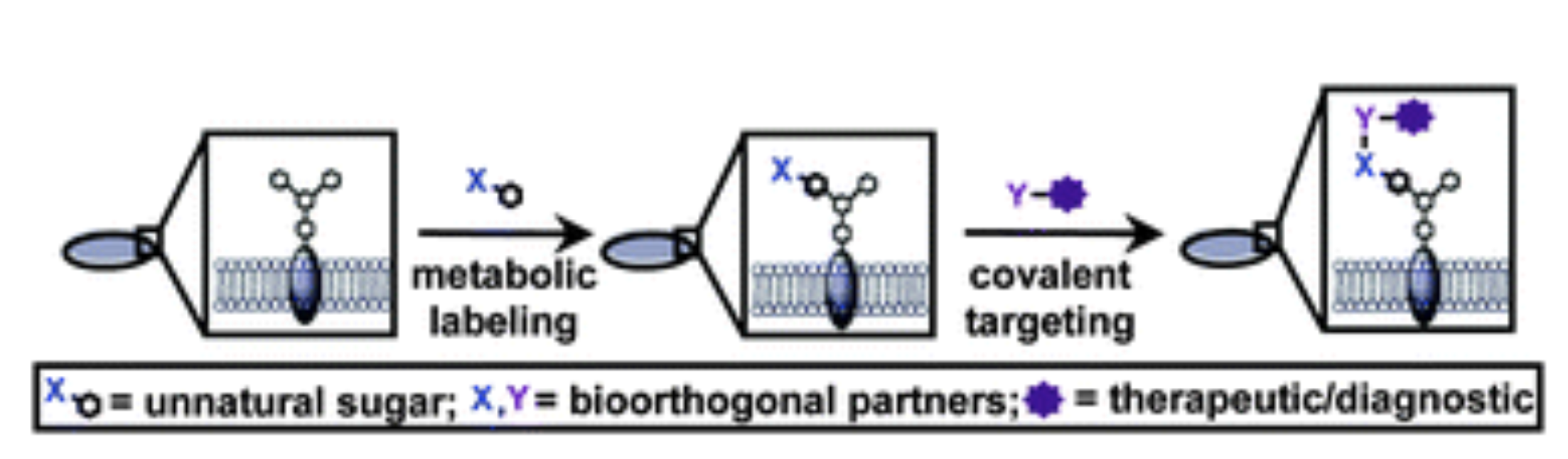
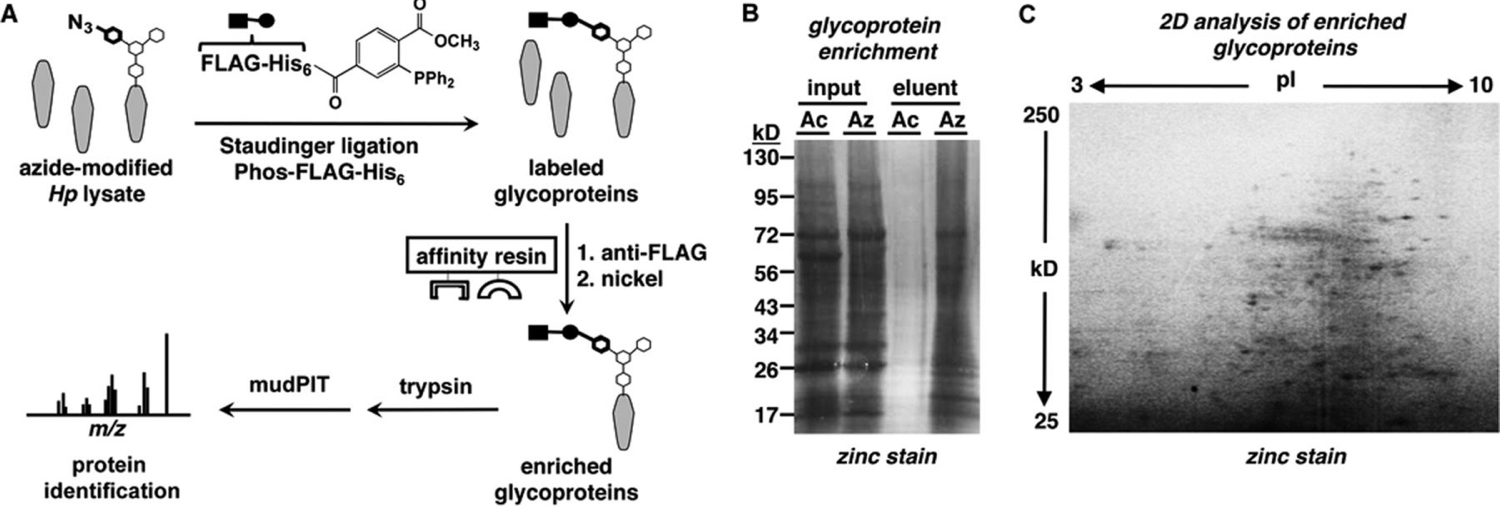
K. Champasa*†, S. A. Longwell*†, A. M. Eldridge, E. A. Stemmler and D. H. Dube, "Targeted identification of glycosylated proteins in the gastric pathogen Helicobacter pylori", Mol. Cell. Proteomics, 2013, 12, 2588-2586.
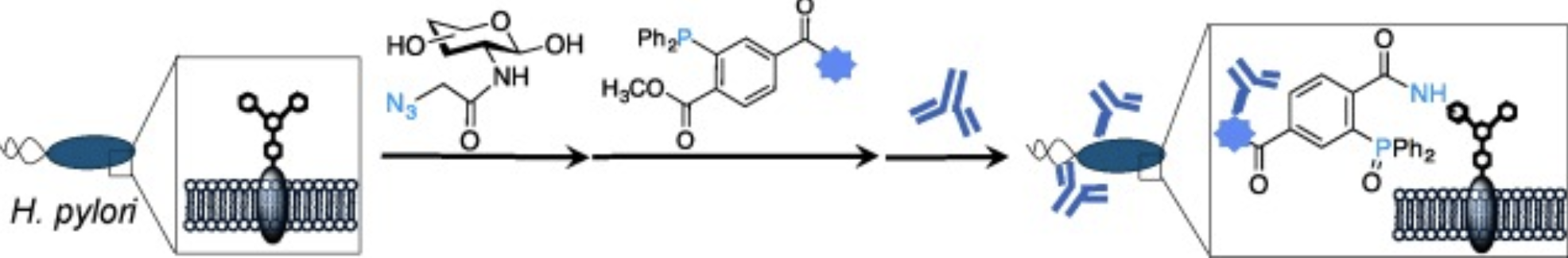
P. Kaewsapsak*, O. Esonu*, and D. H. Dube, "Recruiting the host's immune system to target Helicobacter pylori's surface glycans", ChemBioChem, 2013, 14, 721-726
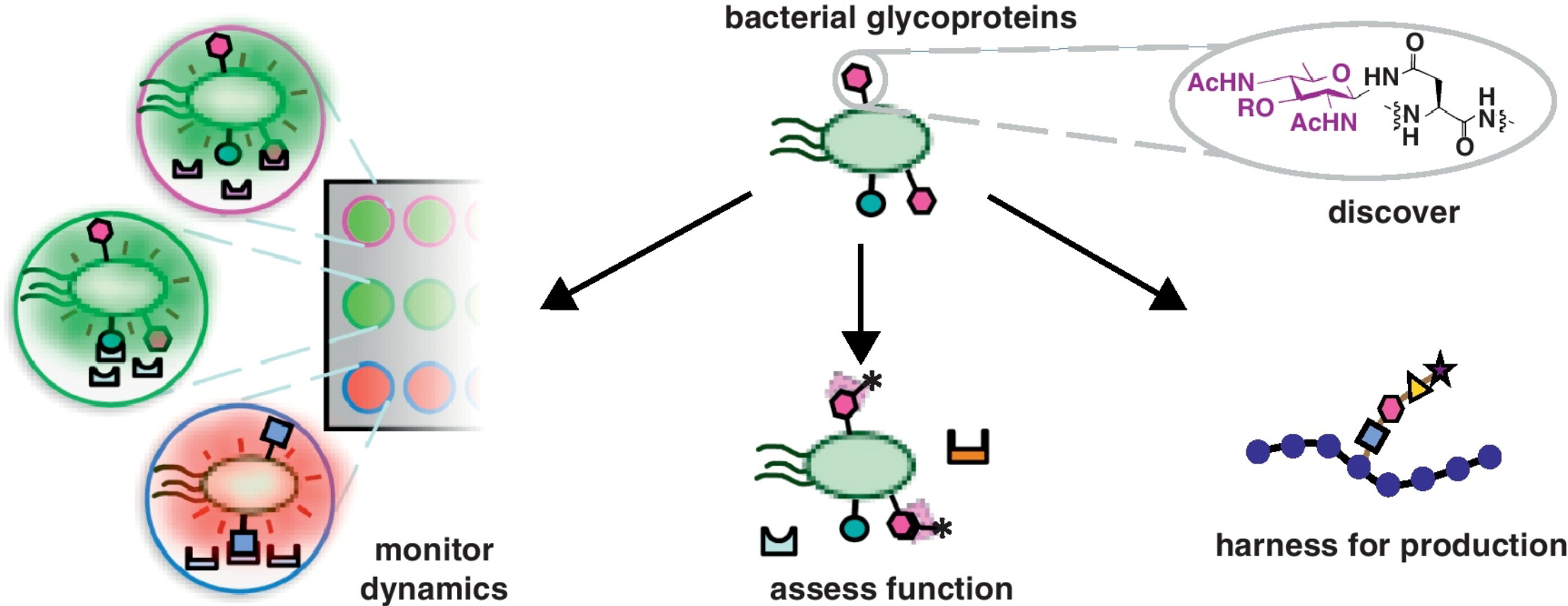
S. A. Longwell* and D. H. Dube, "Deciphering the bacterial glycocode: recent advances in bacterial glycoproteomics", Curr. Opin. Chem. Biol., 2013, 17, 41-48
D. H. Dube. "Metabolic labeling of bacterial glycans with chemical reporters." In Bacterial glycomics: Current research, technology, and applications. Twine, S. M., ed. Horizon Scientific Press, Norfolk (UK), 2012
D. H. Dube†, B. Li†, E. J. Greenblatt, S. Nimer, A. K. Raymond* and J. J. Kohler, "A two-hybrid assay to study interactions within the secretory pathway", PLOS One, 2010, 5, e15648. [† These authors contributed equally to this work.]
D. H. Dube, K. Champasa* and B. Wang*, "Chemical tools to discover and target bacterial glycoproteins",Chem. Commun., 2011, 47, 87-101.
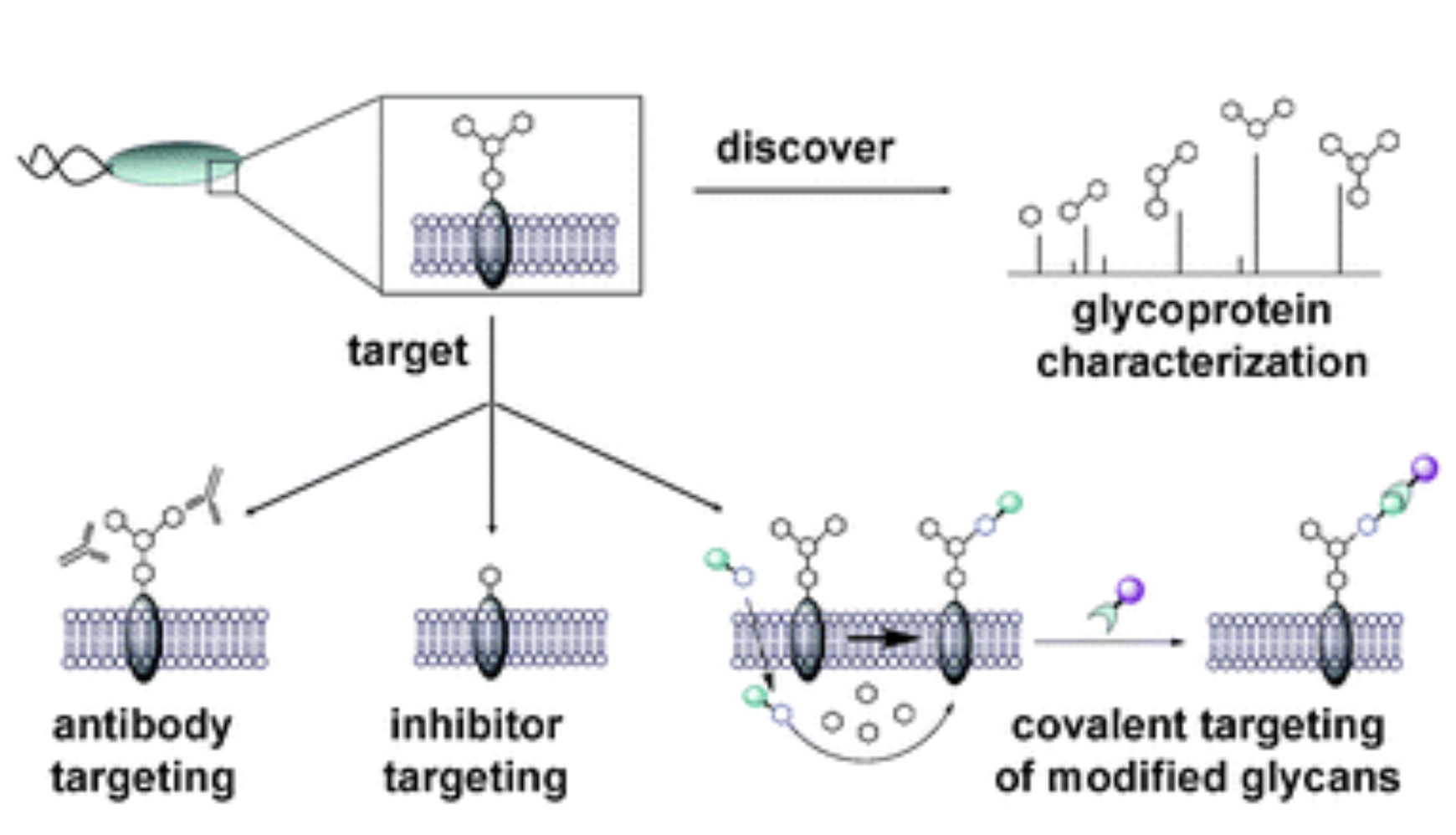
P. V. Chang, D. H. Dube and C. R. Bertozzi, "A strategy for the selective imaging of glycans using caged metabolic precursors", J. Am. Chem. Soc., 2010, 132, 9516-9518.
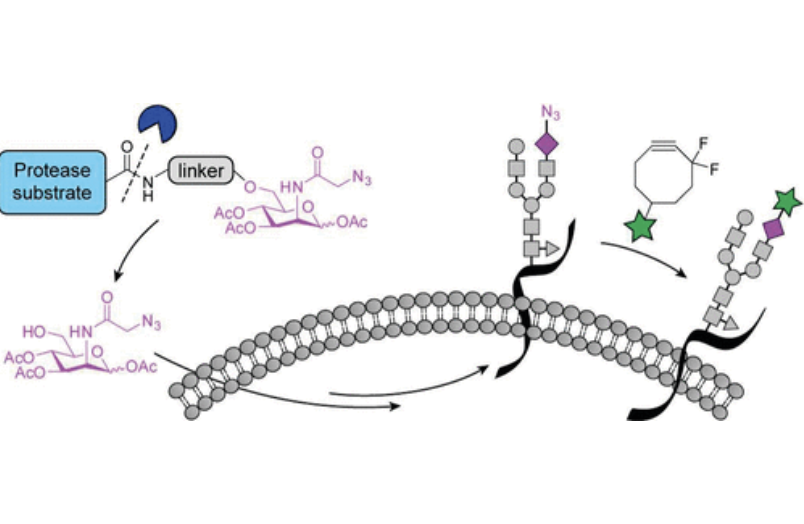
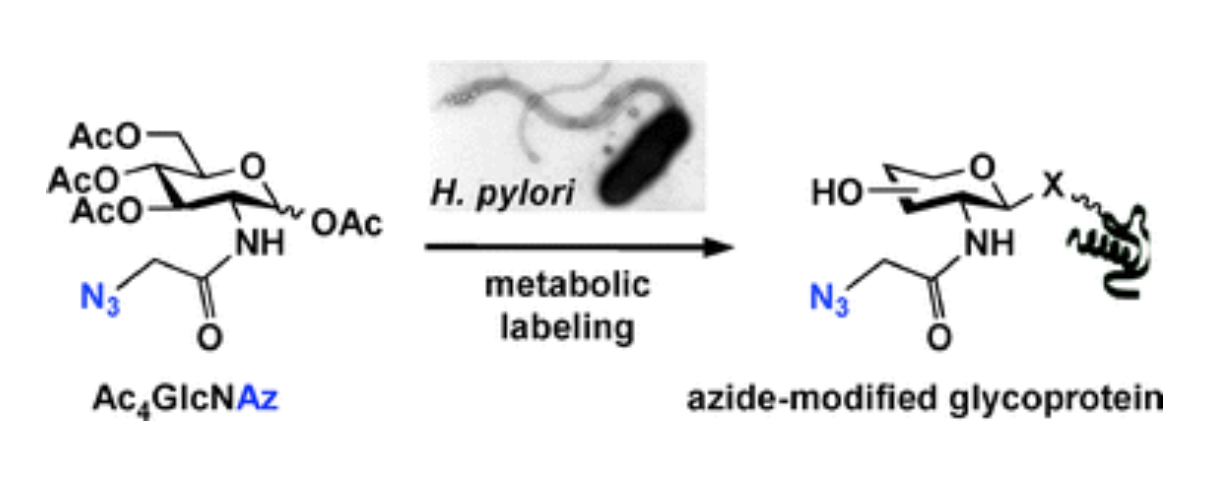
M. B. Koenigs*, E. A. Richardson* and D. H. Dube, "Metabolic profiling of Helicobacter pylori glycosylation",Mol. BioSyst., 2009, 5, 909-912.
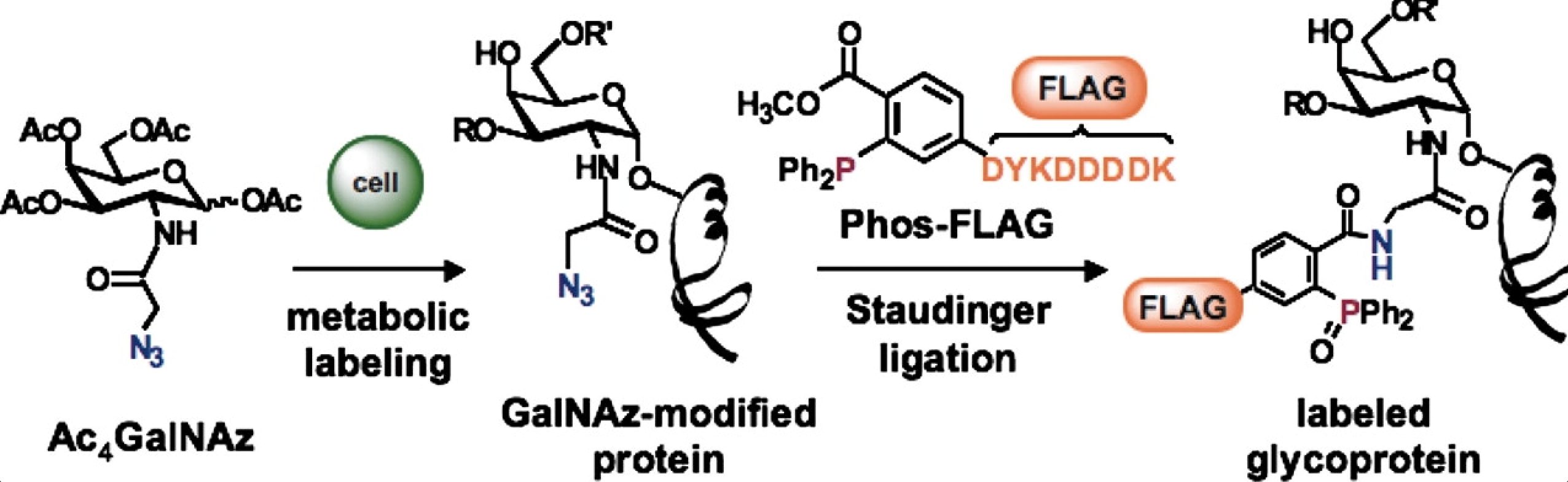
D. H. Dube, C. L. De Graffenried and J. J. Kohler, “Regulating cell surface glycosylation with a small-molecule switch”, Meth. Enzymol., 2006, 415, 213-229.
D. H. Dube†, J. A. Prescher†, C. N. Quang* and C. R. Bertozzi, “Probing mucin-type O-linked glycosylation in living animals”, Proc. Nat’l Acad. Sci. USA, 2006, 103, 4819-4824. [†These authors contributed equally to this work.]
D. H. Dube and C. R. Bertozzi, “Glycans in cancer and inflammation: potential for therapeutics and diagnostics”, Nat. Rev. Drug Disc., 2005, 4, 477-488.
J. A. Prescher†, D. H. Dube†, and C. R. Bertozzi, “Chemical remodeling of cell surfaces in living animals”Nature, 2004, 430, 873-877. [†These authors contributed equally to this work.]
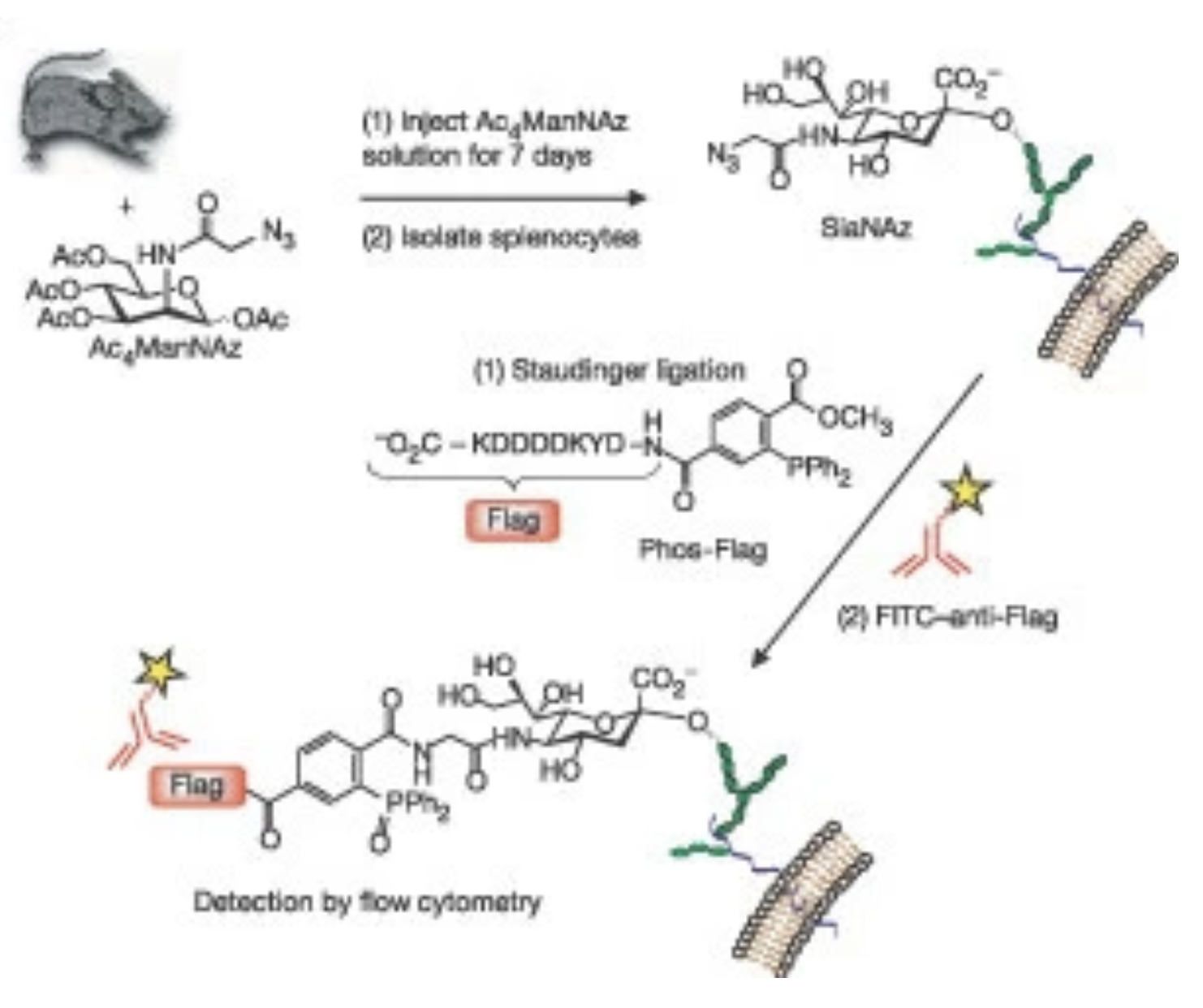
D. H. Dube and C. R. Bertozzi, “Metabolic oligosaccharide engineering as a tool for glycobiology”, Curr. Opin. Chem. Biol., 2003, 7, 1-10.
S. J. Luchansky, H. C. Hang, E. Saxon, J. R. Grunwell, C. Yu, D. H. Dube and C. R. Bertozzi, “Constructing azide-labeled cell surfaces using polysaccharide biosynthetic pathways”, Meth. Enzymol., 2003, 362, 249-272.
*indicates undergraduate co-author, †These authors contributed equally to this work.
































